DNA Methylation and Stress Genes
Compound evidence detail2 SCRs / 4 parts
- Documentedthe canine methylation pattern reported by Awalt 2024 for NR3C1 and OXTR in dogs with measured early-life history
- Heuristicthe JB extrapolation that ordinary calm raising produces durable epigenetic advantages in dogs beyond preventing adverse environments
- Documentedthe Awalt 2024 finding that NR3C1 and OXTR methylation in dogs covaries with documented early-life experience
- Heuristicthe JB inference that those documented methylation differences in dogs predict specific later health or developmental outcomes
DNA methylation is the best-studied epigenetic mechanism in stress biology. It usually refers to the addition of a methyl group to cytosine at CpG sites, often in regulatory regions that influence whether a gene is read more or less actively. In stress research, methylation matters because it offers a plausible mechanism by which early environment can leave lasting effects on how stress-regulation genes are expressed. Documented
What It Means
Methylation does not change the genetic code itself. It changes how accessible a region of DNA is to the cellular machinery that reads it. The safe general rule is that more methylation near key regulatory regions often means less transcription, while less methylation often means greater transcriptional accessibility. That rule is not absolute for every gene in every tissue, but it is the right starting point for stress-gene literature. Documented
Why Early Life Matters So Much
Development is when methylation patterns are especially plastic. Early life is not the only period when methylation can shift, but it is one of the periods when environment appears to have especially strong leverage. That is why developmental stress science keeps returning to caregiving, adversity, predictability, and chronic background conditions rather than only to dramatic isolated events. The question is not only whether the organism was stressed once. The question is whether the developmental environment helped set a durable regulatory pattern.
The Main Stress-Related Target Genes
Several genes appear repeatedly in this literature: NR3C1 (the glucocorticoid receptor gene), FKBP5 (which influences glucocorticoid signaling dynamics), BDNF (involved in plasticity and neuronal support), and OXTR (the oxytocin receptor gene). Documented The dog literature in this dispatch is strongest for NR3C1 and OXTR. The broader cross-species literature is richer across the larger set.
Why It Matters for Your Dog
The strongest mechanistic story comes from rodent maternal-care work and later human stress studies. The general pattern is that early adversity or low-quality caregiving can be associated with methylation differences in genes relevant to stress regulation, and those methylation patterns can track later differences in reactivity or recovery. Documented This does not mean every methylation difference is destiny or that every stress outcome has a single-gene explanation. It means the stress system has a documented molecular regulatory layer.
SCR-094 is the central dog entry. Awalt and colleagues found that dogs with more adverse early-life histories differed in methylation on NR3C1 and OXTR. Those differences were associated with cortisol reactivity and attachment-related outcomes. Documented That is enough to support a serious canine claim: stress- and bonding-related genes in dogs show methylation patterns that vary with early-life conditions. It is not enough to support a fully detailed canine map for every major stress gene. The dog evidence is still smaller, more associative, and less experimentally controlled than the classic rodent literature.
What This Page Does Not Claim
Because DNA methylation is so mechanistically attractive, it is easy to overread it. This page does not claim that every early experience leaves a stable methylation signature, every methylation difference has a large behavioral consequence, or specific family practices have already been shown to create one predictable canine methylation profile. The strongest takeaway is more modest and more defensible: early-life context in dogs can be associated with methylation differences in genes relevant to stress and bonding. Documented
The calmness layer often argues that chronic background conditions matter more than dramatic moments. Methylation science helps explain why that is biologically plausible: low-level developmental conditions can shape which stress-related genes are more or less readily expressed.
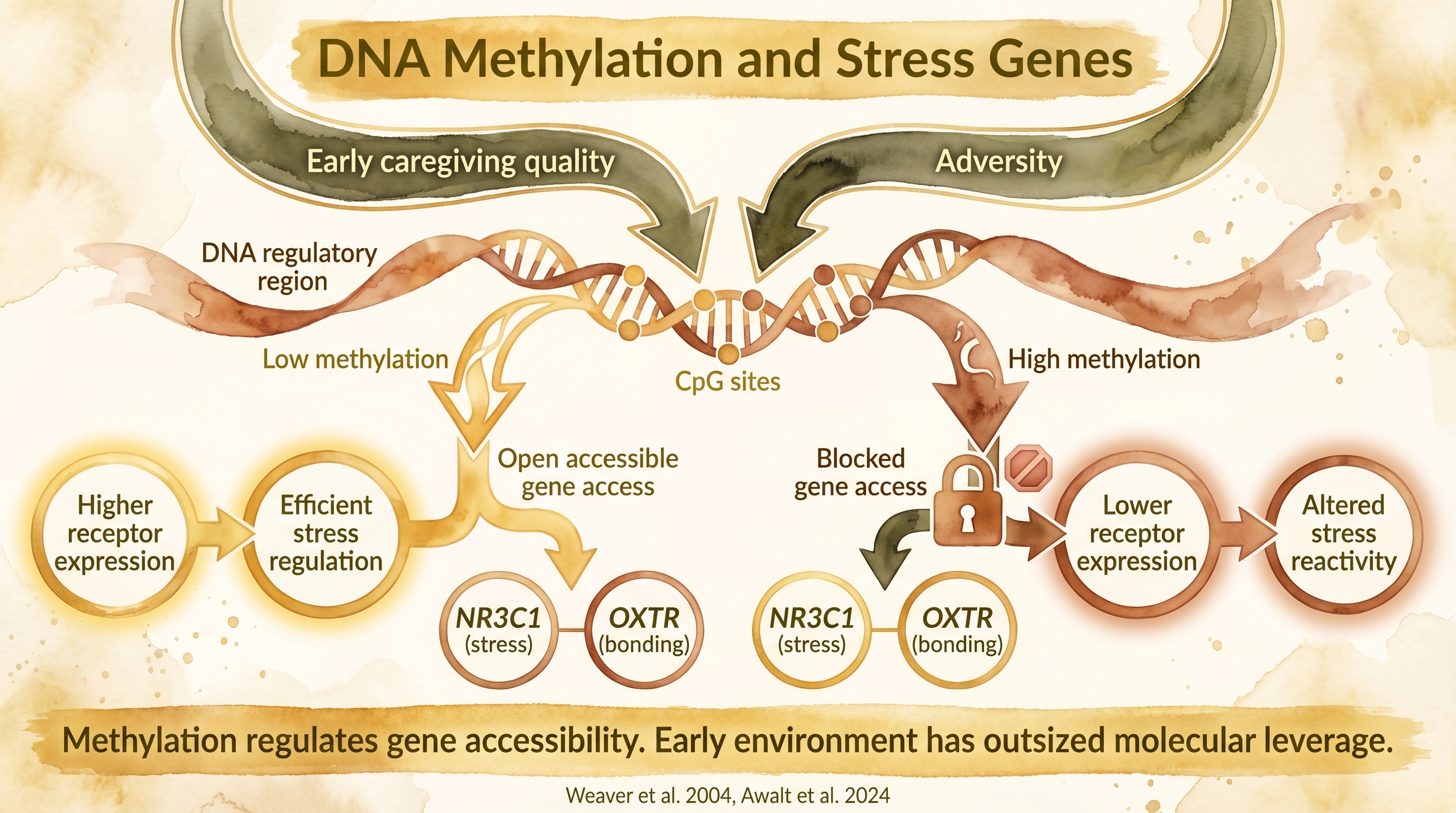
Early caregiving quality physically marks stress-response genes through methylation, altering lifelong cortisol regulation.
Key Takeaways
- DNA methylation is the best-studied epigenetic mechanism in stress biology.
- Early development is a particularly sensitive period for methylation patterning.
- The broader mammalian literature implicates several stress-related genes, especially NR3C1.
- In dogs, the strongest direct evidence currently centers on NR3C1 and OXTR rather than a fully mapped stress-gene catalog.
The Evidence
- Weaver, I. C. G. et al. (2004)rats
Linked maternal care differences with methylation changes in glucocorticoid-related regulatory regions. - McGowan, P. O. et al. (2009)humans
Extended the stress-methylation link to human tissue in the context of childhood abuse history. - Szyf, M. and related mammalian synthesismultiple mammals
Positioned DNA methylation as a central mechanism in environmental programming of stress-related gene expression.
- Awalt, S. L. et al. (2024)domestic dogs
Reported early-life-history associations with methylation differences in NR3C1 and OXTR in dogs. - Stress-immunity source synthesisdomestic dogs
Supports the canine stress-regulation system as epigenetically responsive while leaving many outcome claims provisional.
No study has measured methylation changes at known stress-gene sites in the same dogs before and after documented behavioral or environmental interventions, leaving the causal chain from methylation to outcome untested.
No published study in dogs has mapped whether different stress types or different developmental adversities produce distinct methylation signatures at NR3C1, FKBP5, or other candidates.
SCR References
Sources
- Awalt, S. L., et al. (2024). A dog's life: Early life histories influence methylation of glucocorticoid (NR3C1) and oxytocin (OXTR) receptor genes, cortisol levels, and attachment styles. Developmental Psychobiology.
- McGowan, P. O., et al. (2009). Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nature Neuroscience, 12(3), 342-348.
- Szyf, M., et al. (2005). Maternal programming of steroid receptor expression and phenotype through DNA methylation in the rat. Frontiers in Neuroendocrinology.
- Weaver, I. C. G., et al.